| The spike protein of SARS-CoV-2 (CoV2) is required for cell entry and is the primary target for vaccine and therapy development. Unveiling molecular-scale mechanisms relevant to the diffusion of viral particles and their encounter with the cell membrane receptor (ACE2) is a daunting task. We report on the gain in nanomechanical stability of the CoV2 spike protein in comparison with SARS-CoV from 2002. This information allows us to describe the dynamical processes of recognition. It confirms that the receptor-binding domain (RBD of only ∼200 amino acids) makes a significant contribution to the mechanical stability of the full spike homotrimer. From a biophysical perspective, the RBD plays a fundamental role as a damping element of the massive virus particle’s motion prior to cell recognition while also facilitating viral attachment, fusion, and entry. Our findings add a novel way to address the development of therapies aimed at destabilizing specific key contacts of the protein spike, which are responsible for the increased nanomechanical stability. | 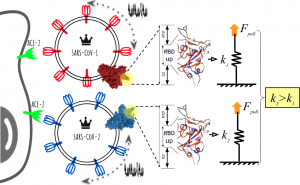 |
Published in:
Featured in an Editor’s Choice web collection focussing on recent breakthroughs in nanobiotechnology. Read the collection here.

